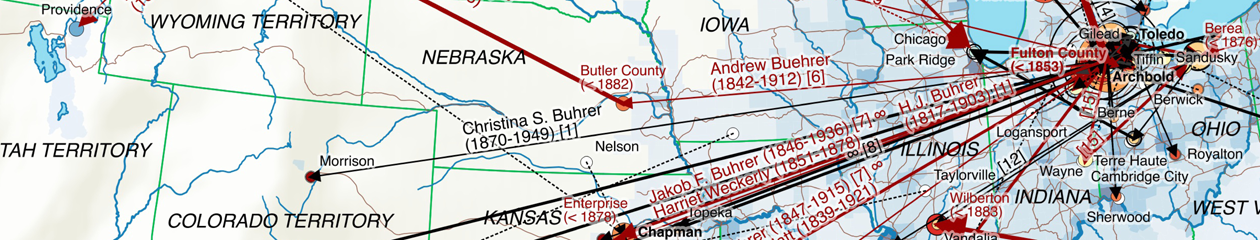
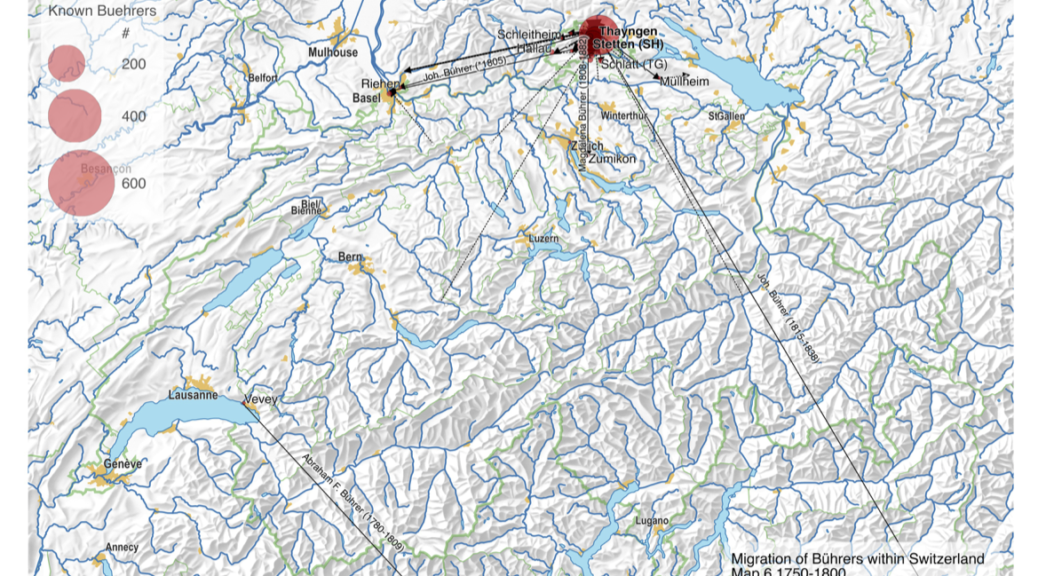
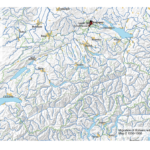
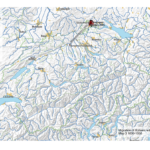
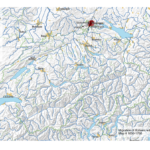
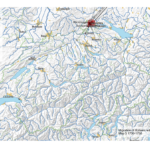
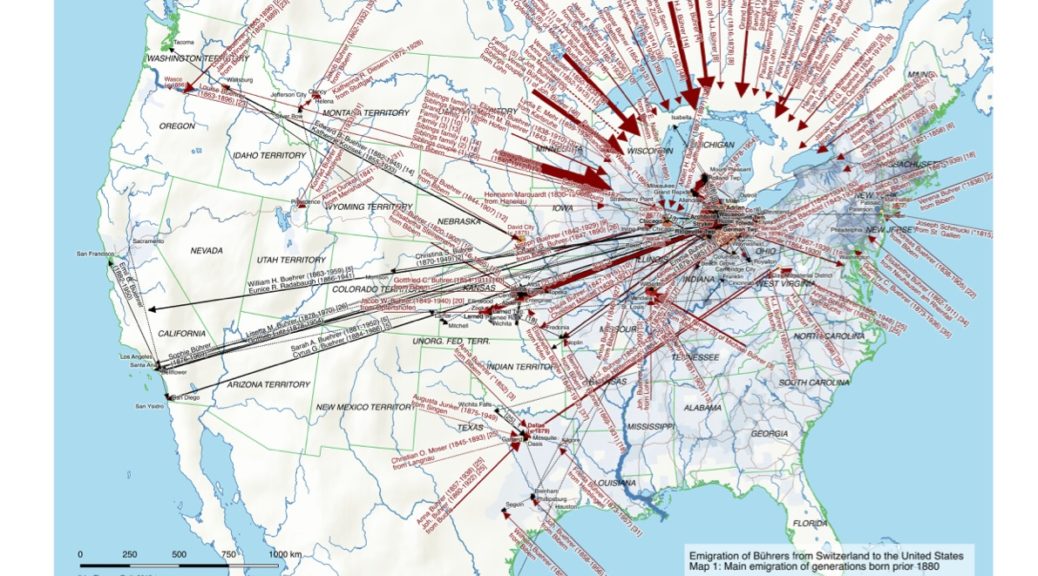
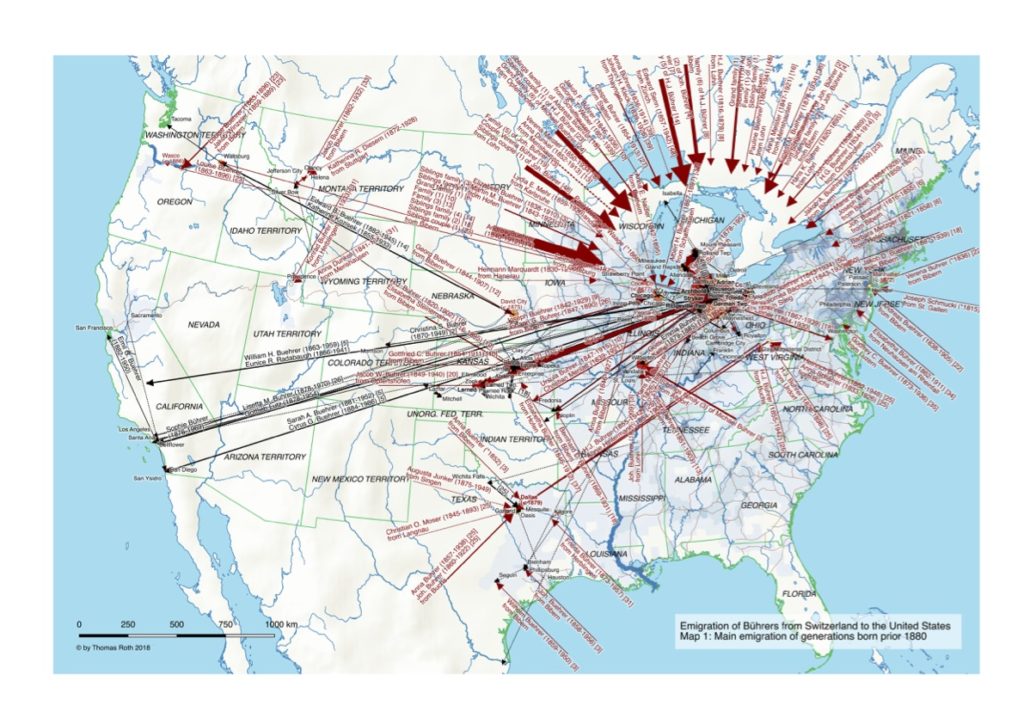
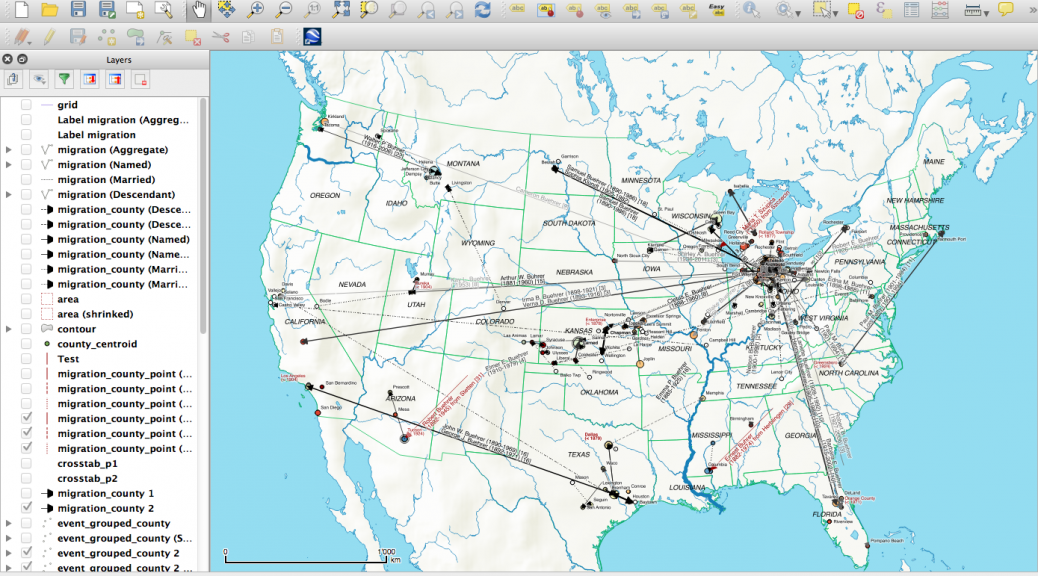
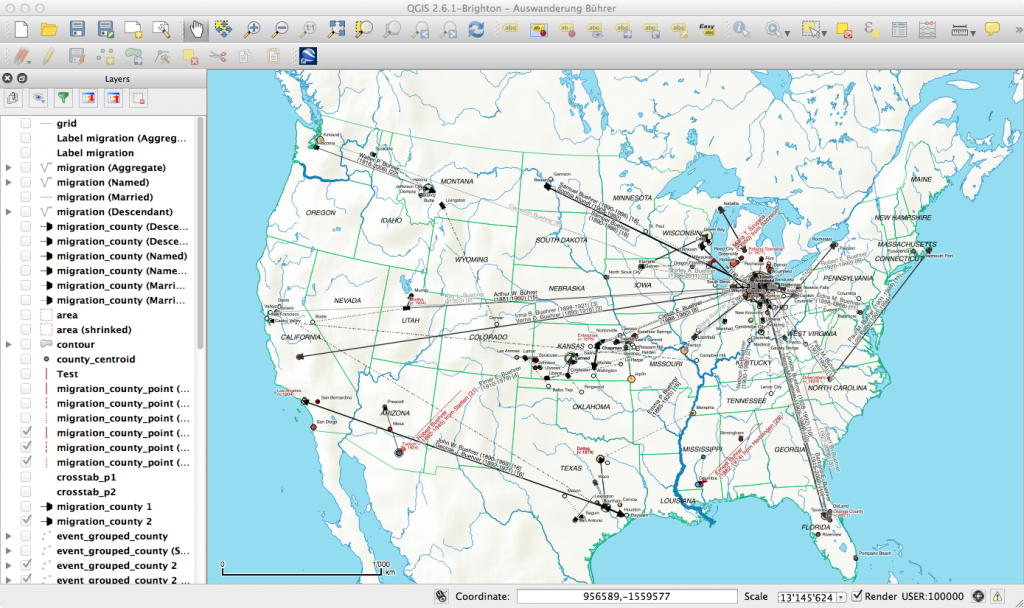
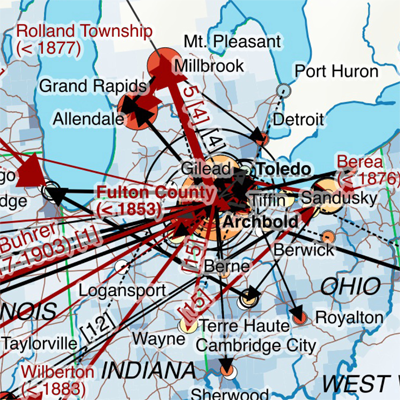
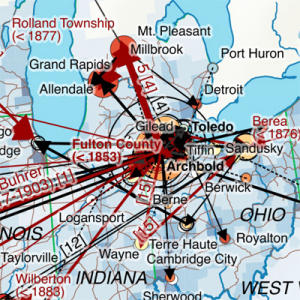
My first map versions used population density and railroads as a proxy for the state and direction of migration. But what about the destination? What was the land cover like at the time of emigration, given the fact that the large majority of emigrating Bührers hauled from the rural countryside and were farmers?
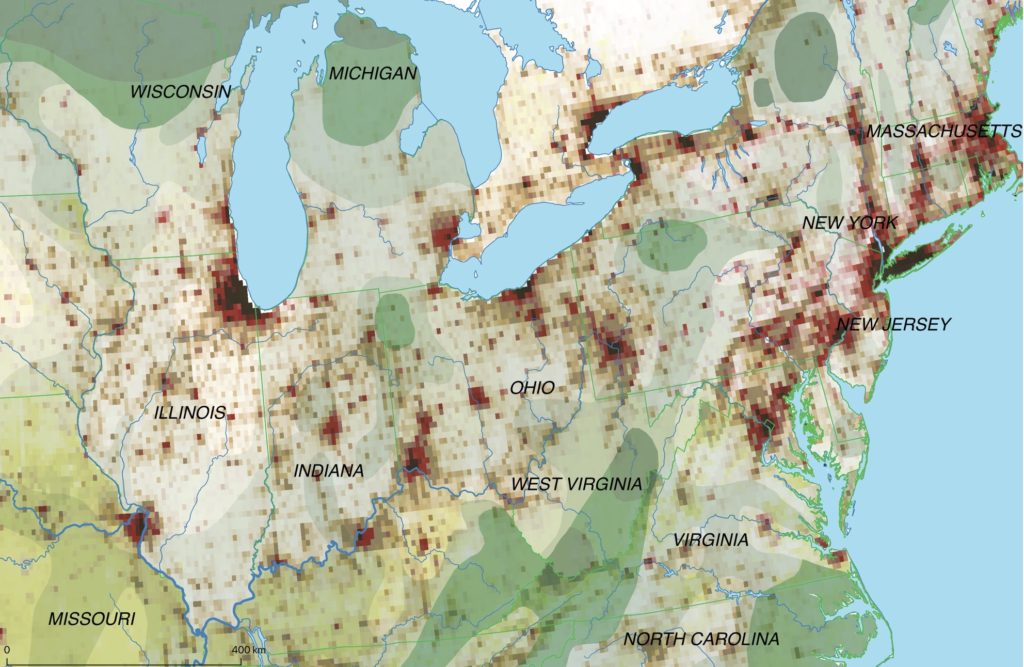
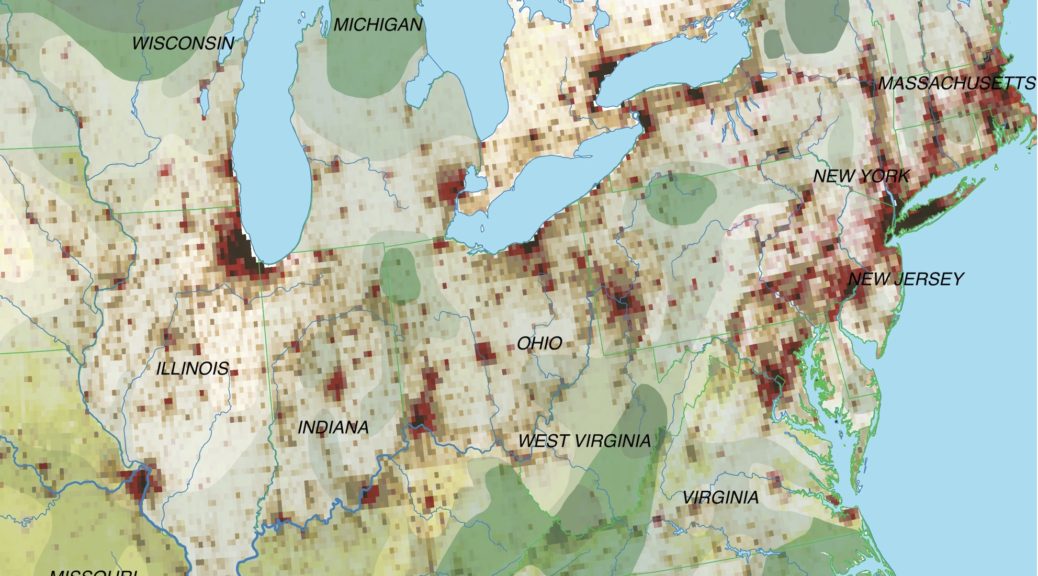
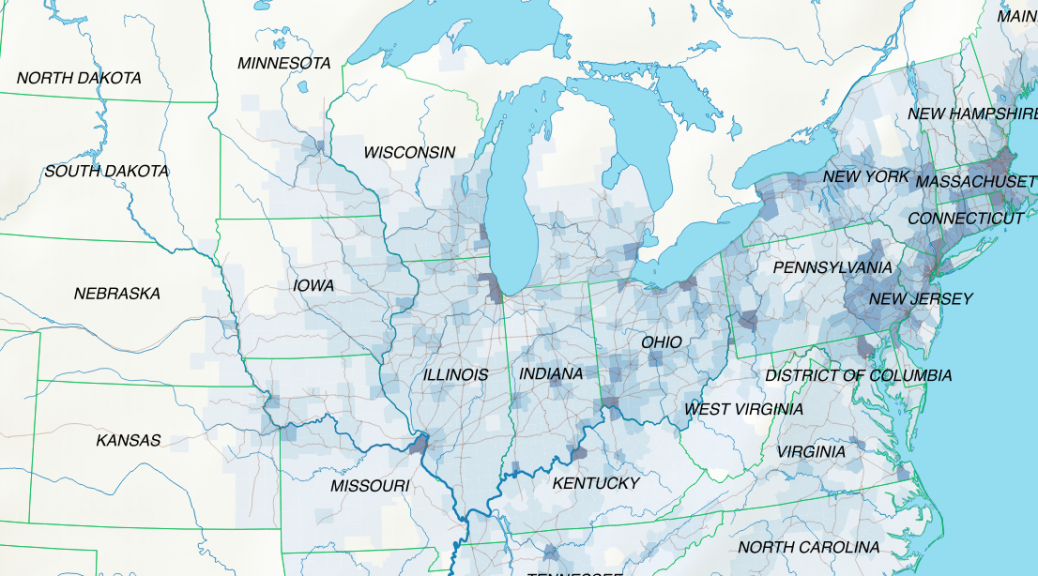
The map above shows the structure of the landcover around 1870. Red indicates urbanised centres, the yellowish green grassland, brown cropland and dark green forestland. Northwestern Ohio was still largely covered by forests (approx. 60% density), with yet only a recognisable patch of cropland around Bryan. Nowadays the forest is – with the exception of small remnants – gone and replaced by cropland. Agriculture completely changed the surface of Ohio.
The sources for this historical land cover were:
- History Database of the Global Environment (HYDE) with historical population, cropland and pasture land use (https://themasites.pbl.nl/tridion/en/themasites/hyde/download/index-2.html)
- Historical woodland density of the conterminous United States, 1873 (https://www.fs.usda.gov/rds/archive/catalog/RDS-2013-0006), a digitised version of a map by Wiliam Brewer
Astonishingly I wasn’t able to find period Ohio maps that include land cover information (e.g. forests).
ISLSCP II Historical Land Cover and Land Use, 1700-1990 provides a source for full land cover (https://daac.ornl.gov/ISLSCP_II/guides/historic_landcover_xdeg.html), albeit only at a very low one-degree resolution that wasn’t useful to produce maps. More interesting is Historical Land-Cover Change and Land-Use Conversions Global Dataset (https://data.nodc.noaa.gov/cgi-bin/iso?id=gov.noaa.ncdc:C00814), with half-degree resolution. There is also data for the Original Natural Vegetation of Ohio (https://apps.ohiodnr.gov/gims/report.asp, see theme id 3135), essentially almost exclusively forestland of different kinds.